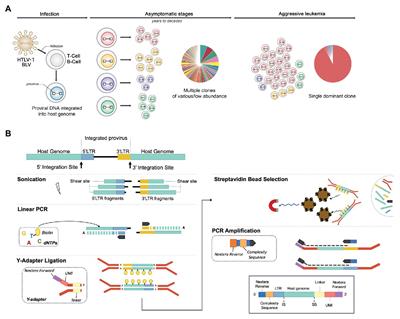
Quantitative Shearing Linear Amplification Polymerase Chain Reaction: An Improved Method for Quantifying Lentiviral Vector Inser

Genome-wide high-throughput integrome analyses by nrLAM-PCR and next-generation sequencing | Nature Protocols

Rapid, accurate mapping of transgene integration in viable rhesus macaque embryos using enhanced-specificity tagmentation-assisted PCR: Molecular Therapy - Methods & Clinical Development

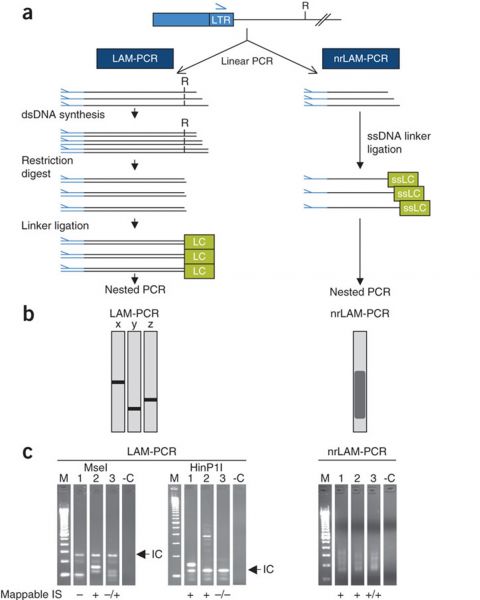
LAM-PCR. a, Outline of the LAM-PCR method. 1, linear PCR by repeated... | Download Scientific Diagram
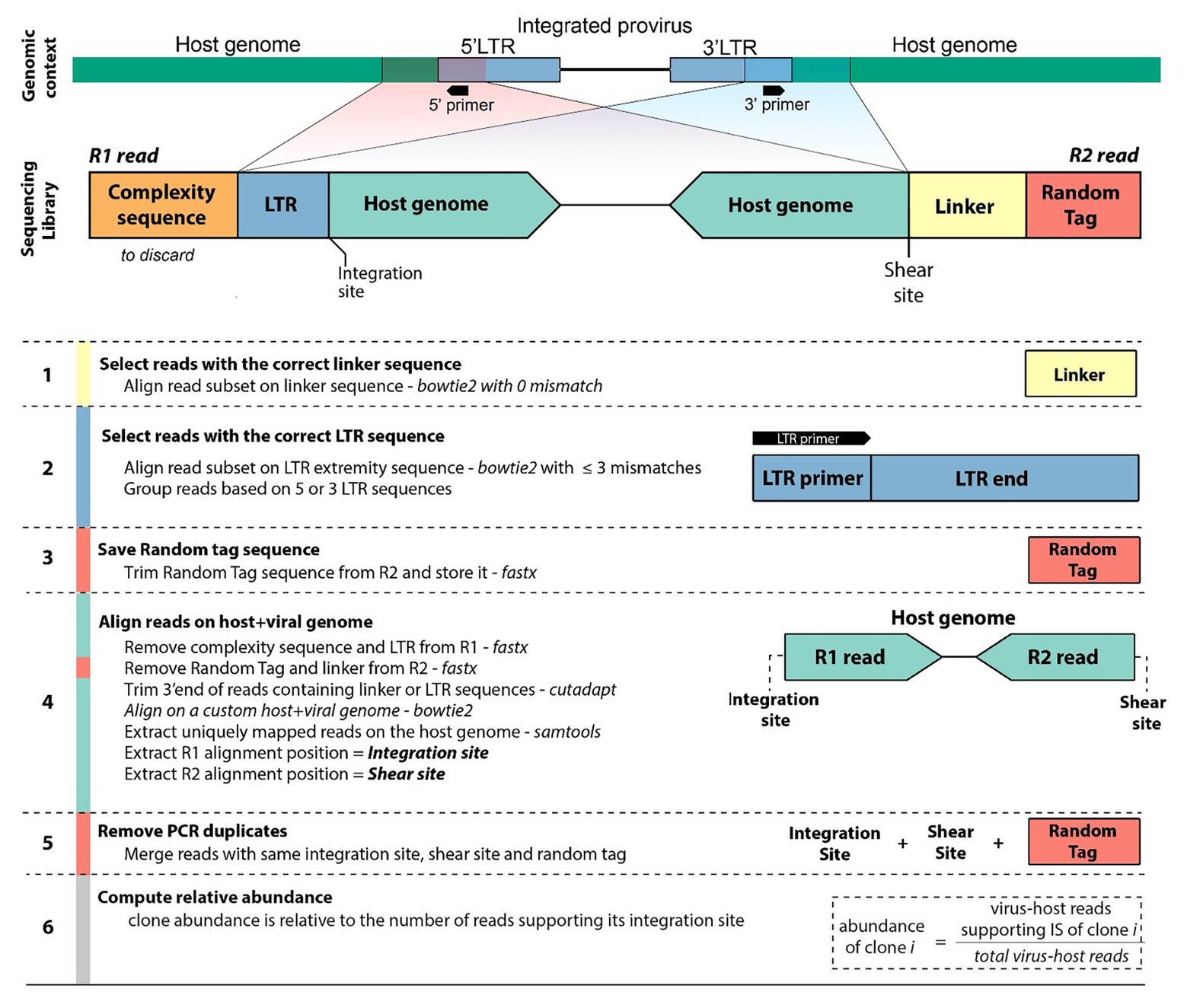
Frontiers | An Improved Sequencing-Based Bioinformatics Pipeline to Track the Distribution and Clonal Architecture of Proviral Integration Sites

Analysis of polyclonal vector integration sites using Nanopore sequencing as a scalable, cost-effective platform | bioRxiv
HaSAPPy: A tool for candidate identification in pooled forward genetic screens of haploid mammalian cells | PLOS Computational Biology

LAM-HGTGTS (Linear Amplification-mediated high-throughput genome-wide translocation sequencing) Our Working Protocol.
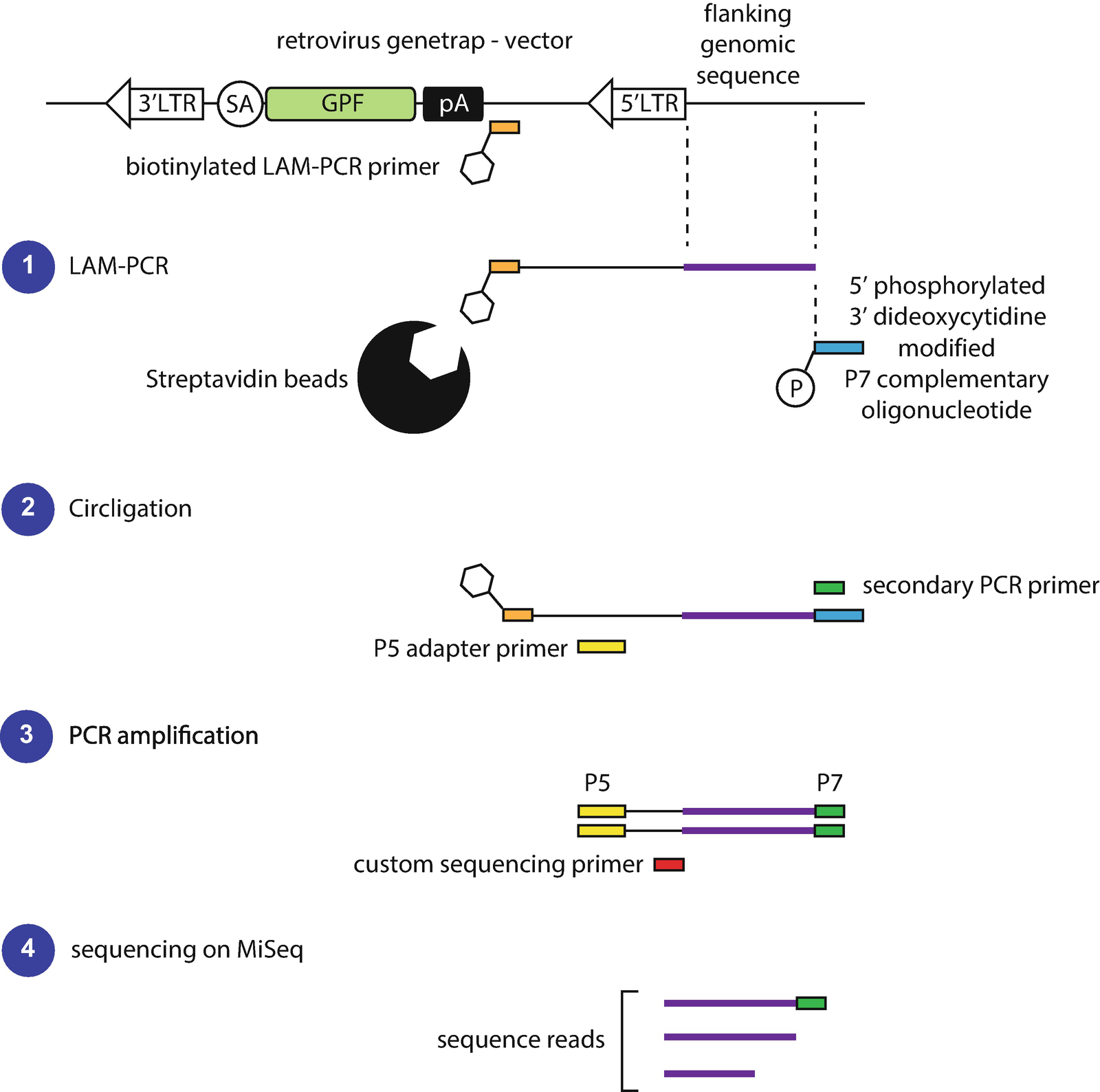
Figure 2 | Screening for Factors Involved in X Chromosome Inactivation Using Haploid ESCs | SpringerLink

High-throughput monitoring of integration site clonality in preclinical and clinical gene therapy studies - ScienceDirect

Schematic of a linear amplification-mediated polymerase chain reaction... | Download Scientific Diagram

DNA sonication inverse PCR for genome scale analysis of uncharacterized flanking sequences - Alquezar‐Planas - 2021 - Methods in Ecology and Evolution - Wiley Online Library

High Efficiency Restriction Enzyme–Free Linear Amplification-Mediated Polymerase Chain Reaction Approach for Tracking Lentiviral Integration Sites Does Not Abrogate Retrieval Bias | Human Gene Therapy